Release Notes for Cytoscape 3.2.0

Cytoscape 3.2.0 is the latest version of Cytoscape 3.x series with new features and bug fixes.
Network loading process is optimized and faster than 3.1.x.
Rendering engine is optimized especially for large networks. Zoom/pan is more efficient and faster than past 3.x series.
Network merge function is parallelized and is much faster for large networks.
Startup process has been optimized and startup time is shorter than 3.1.
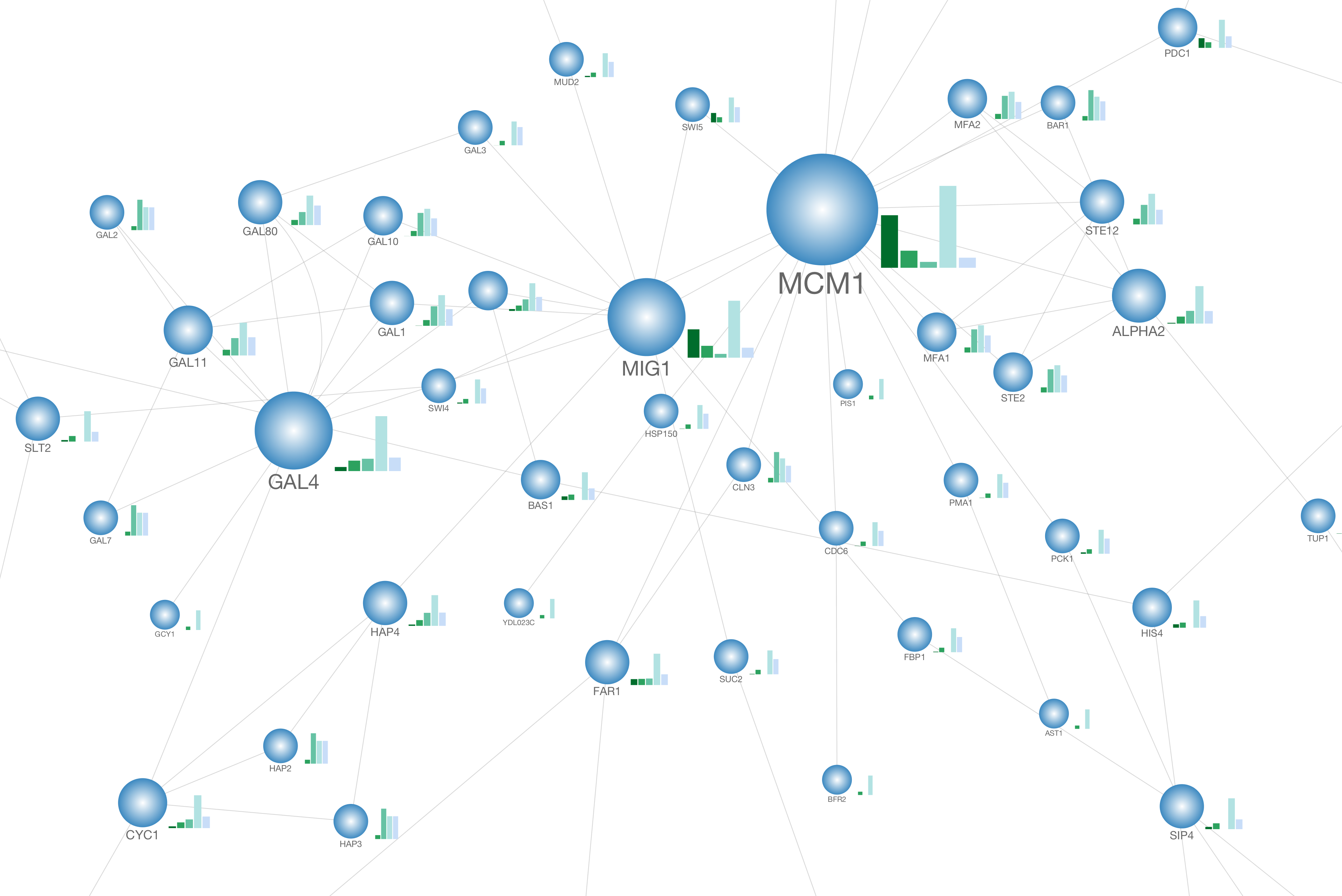
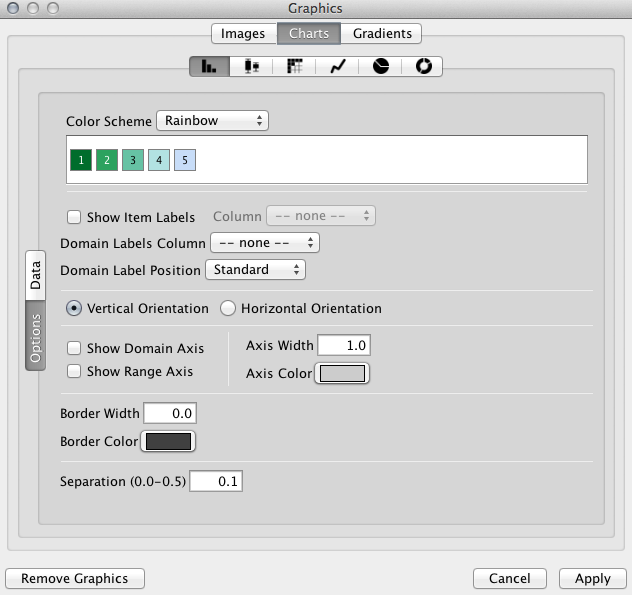
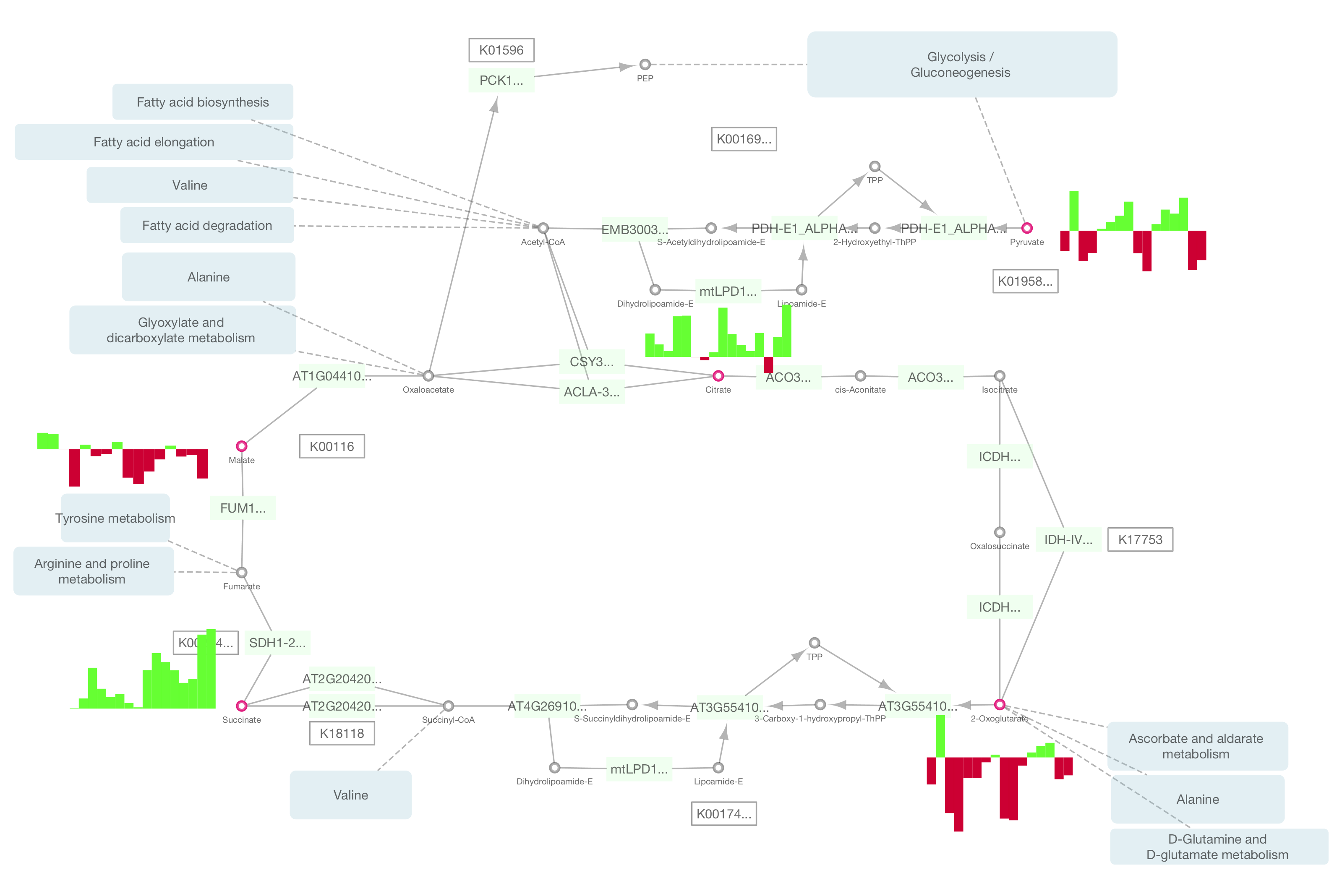
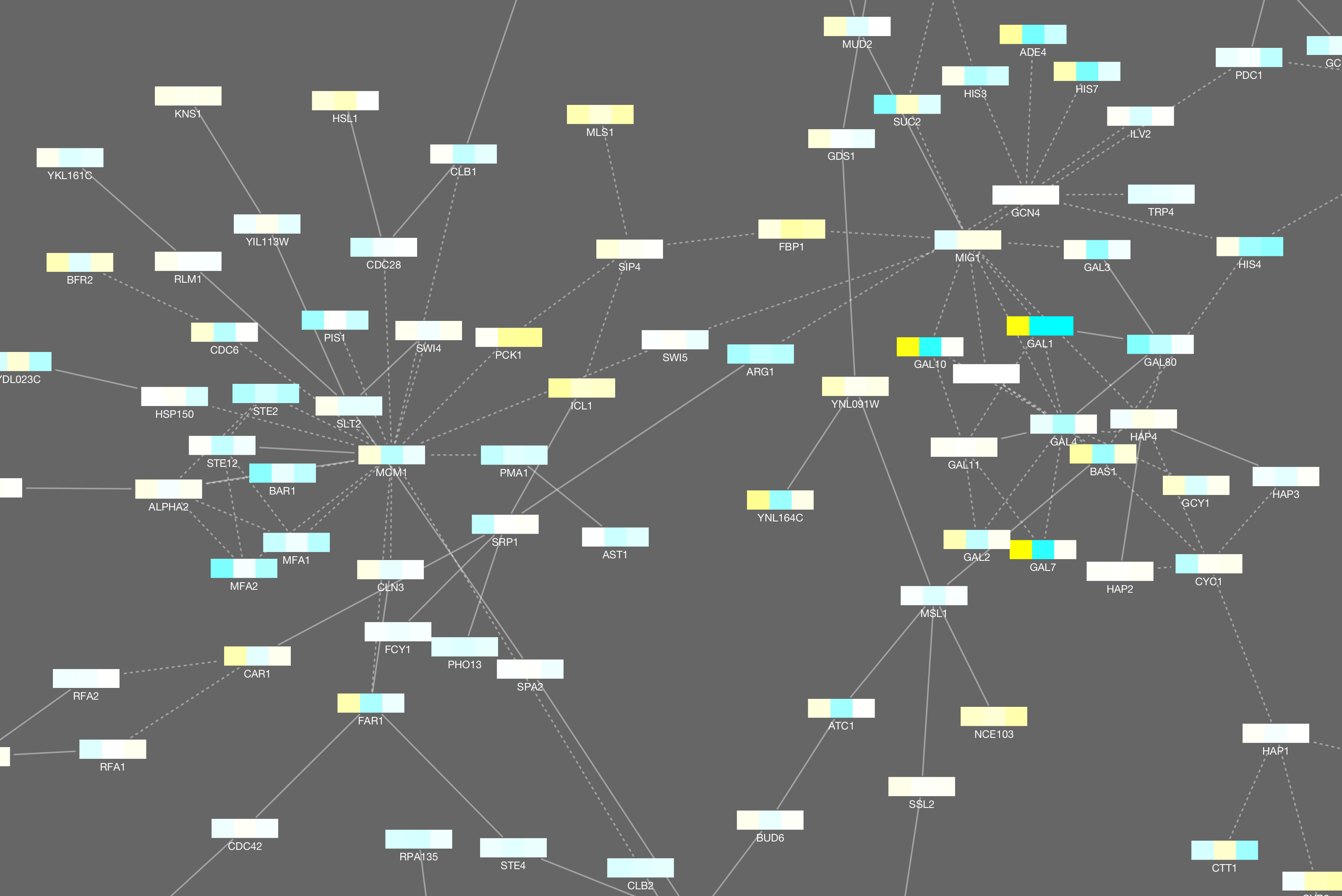
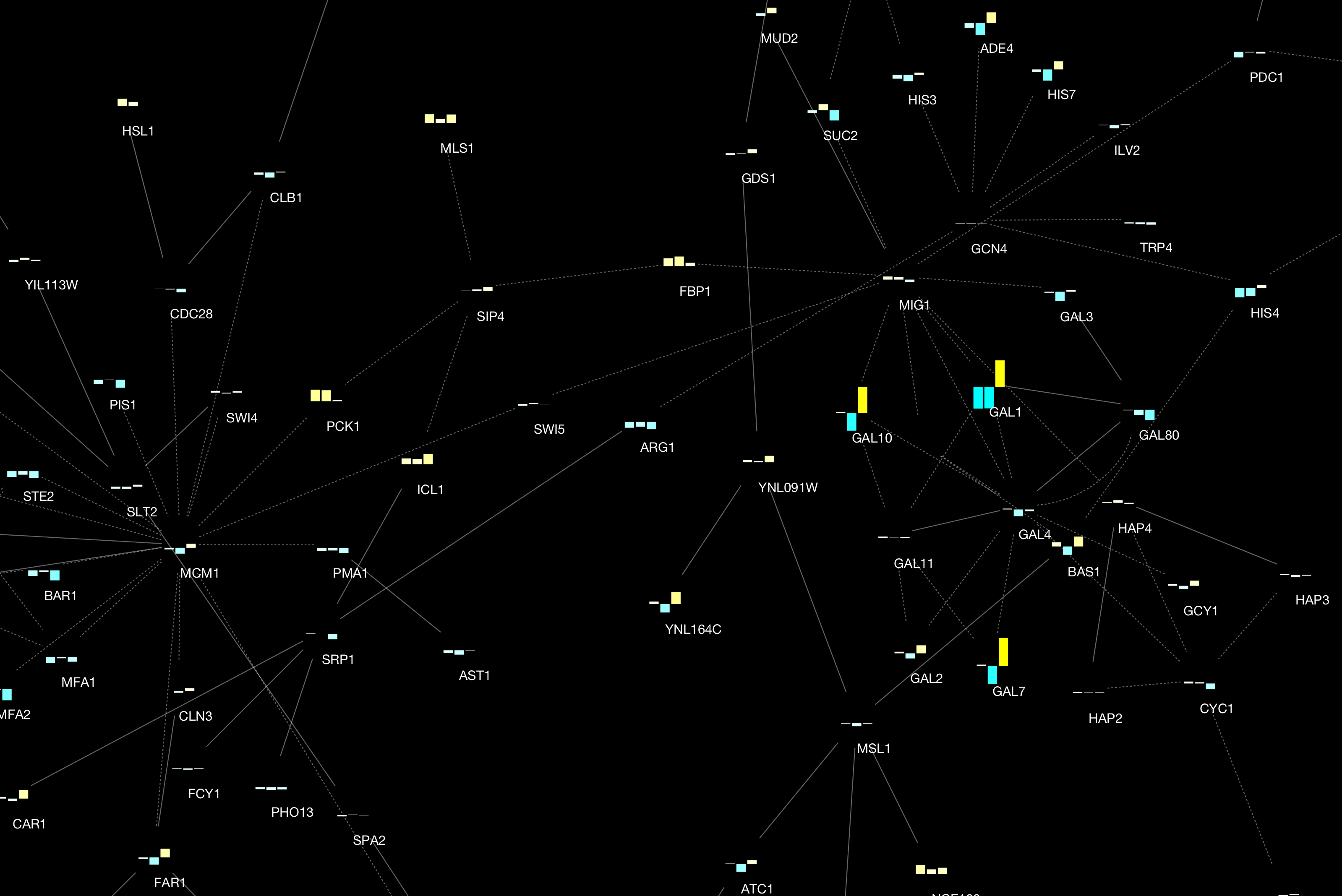
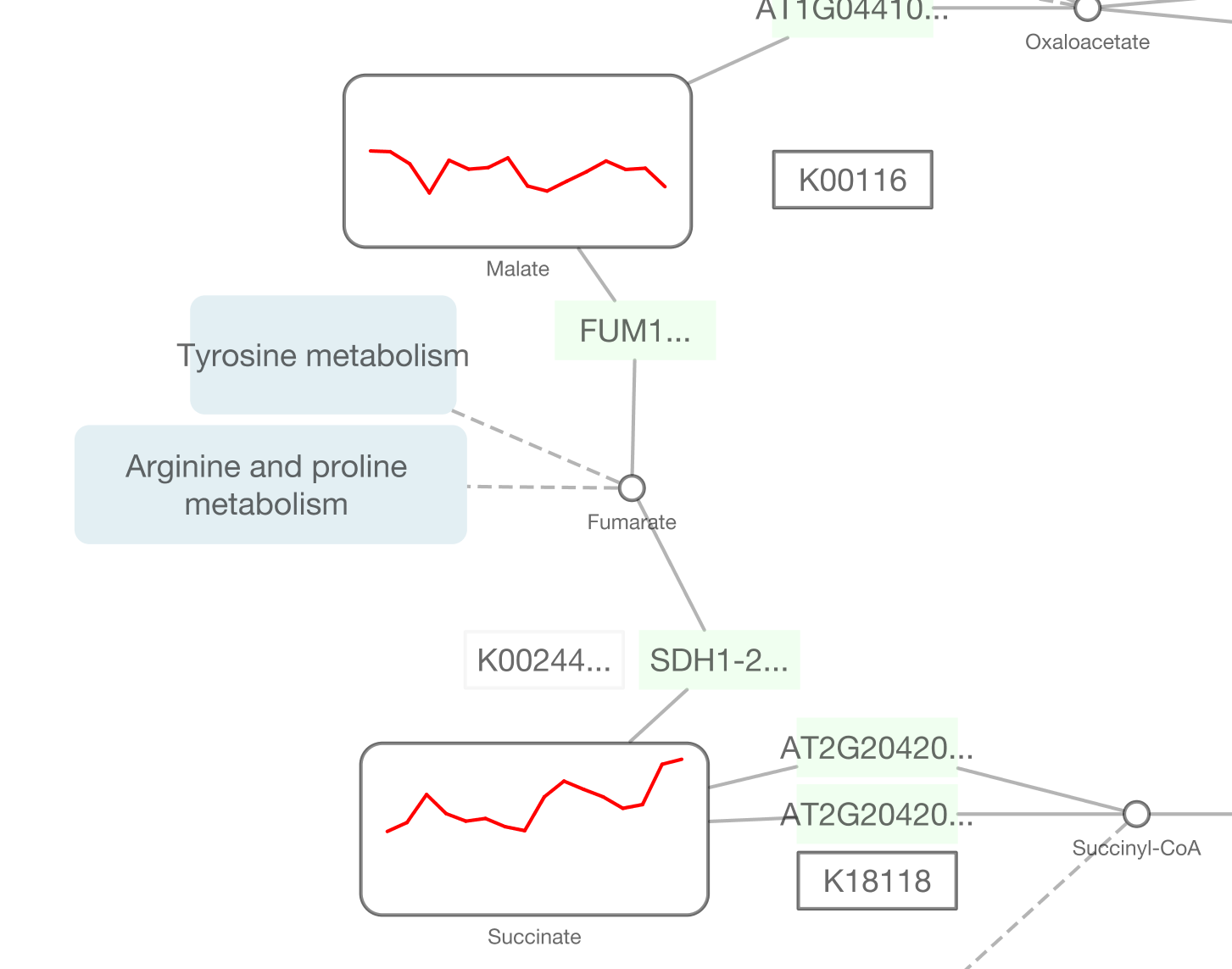
Cytoscape 3.2 has a built-in chart editor. You can use node table data as the source of your charts and render them as high-quality vector graphics. It supports standard charts including: bar, box, line, pie, heatmap, and ring. In addition to charts, it supports gradients.
It is a part of Visual Style editor and you can save everything in your session file.

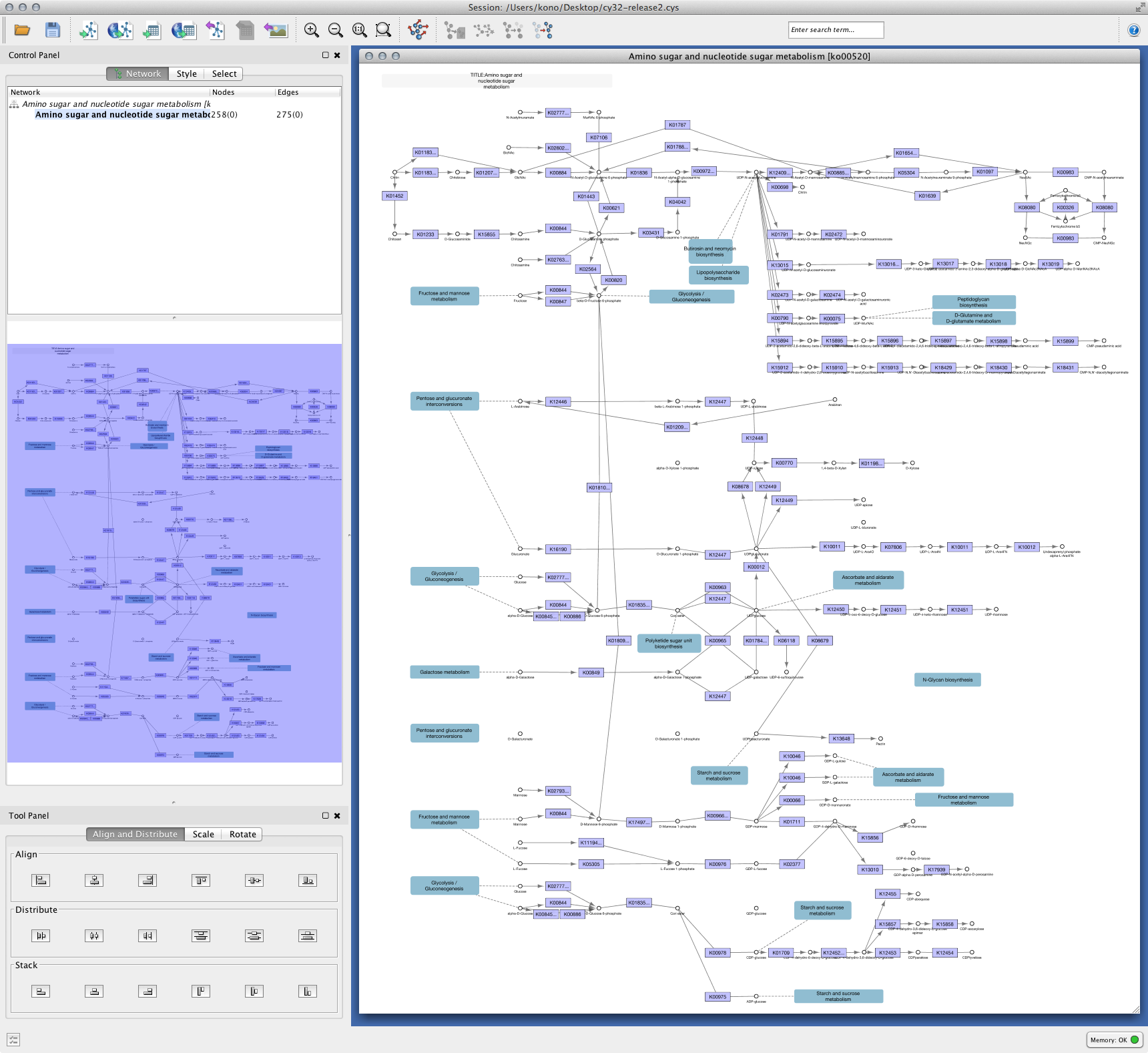
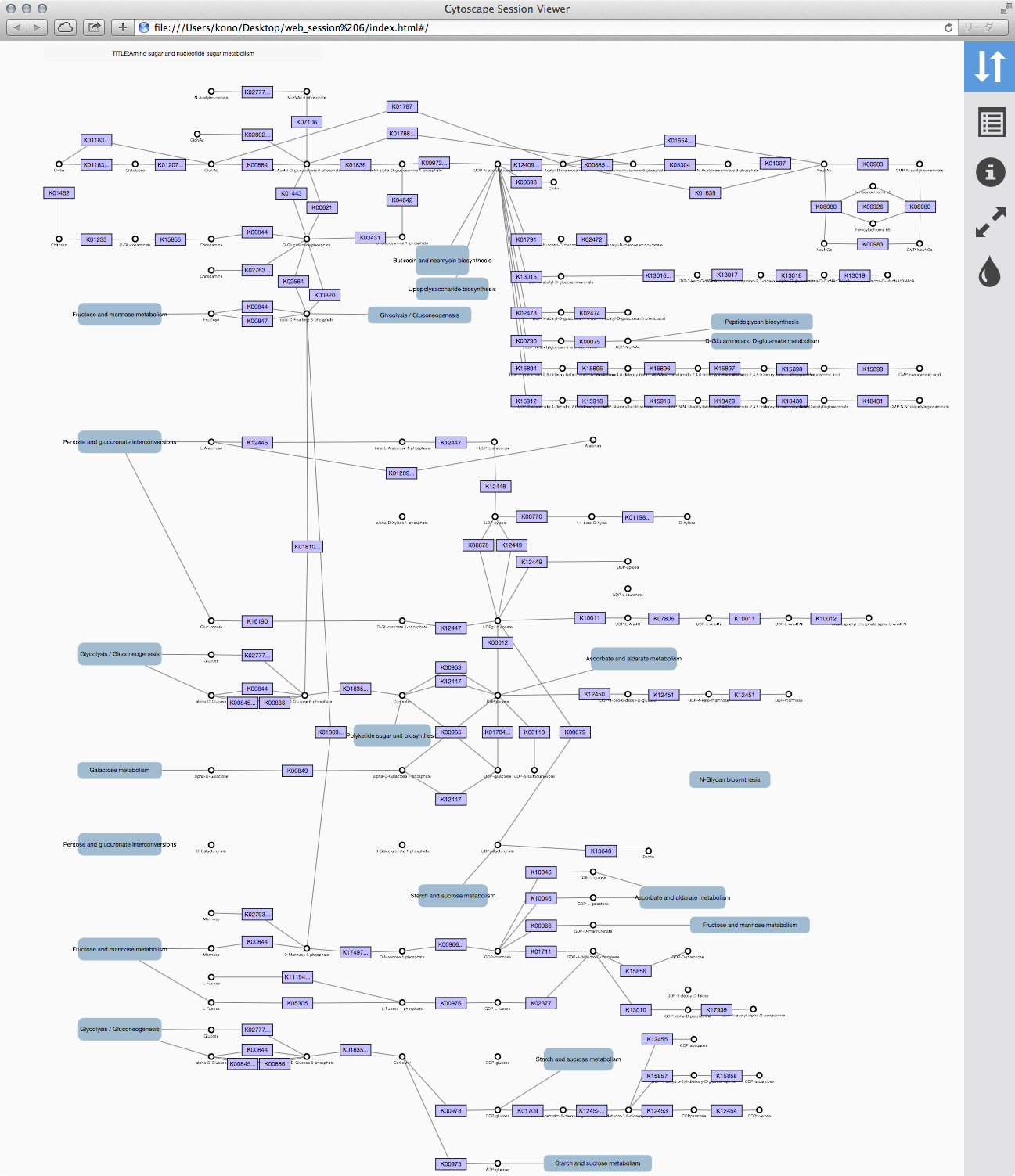
Cytoscape exports your session as Cytoscape.js-based web visualization. You can publish your networks as a complete web application. This feature is for Mac and Linux only. Support for Windows will be available in 3.2.1.
API for Tunables and Commands are cleaned up and more stable (for developers).
While v3.2 continues to support 32 bit OSs, it no longer supports Java 6. We plan to support 32 bit OSs as long as we can, but will eventually drop this support. Desupporting Java 6 matches current industry practice, and allows us to focus our resources on more advanced feature development.
Cytoscape 3.2.0 may run on Java 8, but its support is still experimental. We recommend using Java 7 until we release the official Java 8 compatible version of Cytoscape.
In Cytoscape 3.2, table identifier columns are no longer treated as Strings by default, which can cause a type mismatch error when using numeric identifiers. Because of this, you will need to manually change the ID column data type to import a table with a numeric identifier into a network using a String column as an identifier (such as “shared name”). To do this, right-click the column heading in the table import preview and change the selected type to String before importing.
For Japanese, Korean, and Chinese users, rendering a network to a PDF file can result in loss of labeling information in the PDF. As a workaround, users can generate any type of image file and use the image file instead.
The latest privacy policy is posted here.
On Linux, on the proxy configuration dialog box, fonts are clipped and messages are truncated.
On all platforms, users installing Cytoscape directly from a ZIP or TAR file should manually clear the Cytoscape cache by deleting the CytoscapeConfiguration folder in the user’s home directory.
On all platforms, all Cytoscape session files recorded since v3.0.1 are encoded in UTF-8 instead of the native language encoding. This makes session files portable between workstations in all locales. Users in Japan, Korea, and China are most affected – existing v3.0.0 or v2.x session files must be translated to UTF-8 using a platform-dependent editor (which most users are already using for this purpose). Users in Europe and the Americas are affected, too, if they use characters beyond the standard ANSI 128 – they can translate to UTF-8 using a platform-dependent editor (e.g., Notepad for Windows).
When loading multiple networks into the same network collection, it is important to import tables only after each of the networks is loaded. Interspersing table imports with network loads may cause some table columns to become inaccessible to some networks.
Users upgrading from previous Cytoscape versions (e.g., 3.0.2) may notice that the Welcome Screen contains only six of the eight organism networks shown in the user manual. To add the two new networks, terminate Cytoscape, remove the biogrid directory in the CytoscapeConfiguration directory, and restart Cytoscape. For more information on the CytoscapeConfiguration directory, see Launching Cytoscape in manual.
Your bug reports are very important to improve quality of future versions of Cytoscape 3. If you notice any problems, please report them from:
Help → Report a bug...
Or, you can directly report it from Report a bug link on the navigation bar.
We need your feedback to improve Cytoscape 3! Please send your questions and comments to our mailing list.